A computational framework for structuring, optimising, and evaluating traditional health knowledge. We apply techniques from pharmaceutical drug discovery, quantitative finance, and molecular informatics to the domain of botanical and traditional formulations.
01
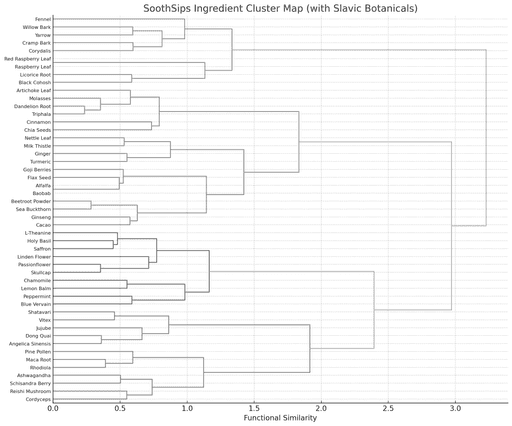
Corpus Structuring
Systematic extraction and normalisation of compound-level data from ethnobotanical literature, clinical databases, and traditional medical texts into queryable knowledge graphs.
02
Receptor-Binding Modelling
Computational docking and affinity scoring of botanical compounds against target receptor panels using validated molecular simulation pipelines.
03
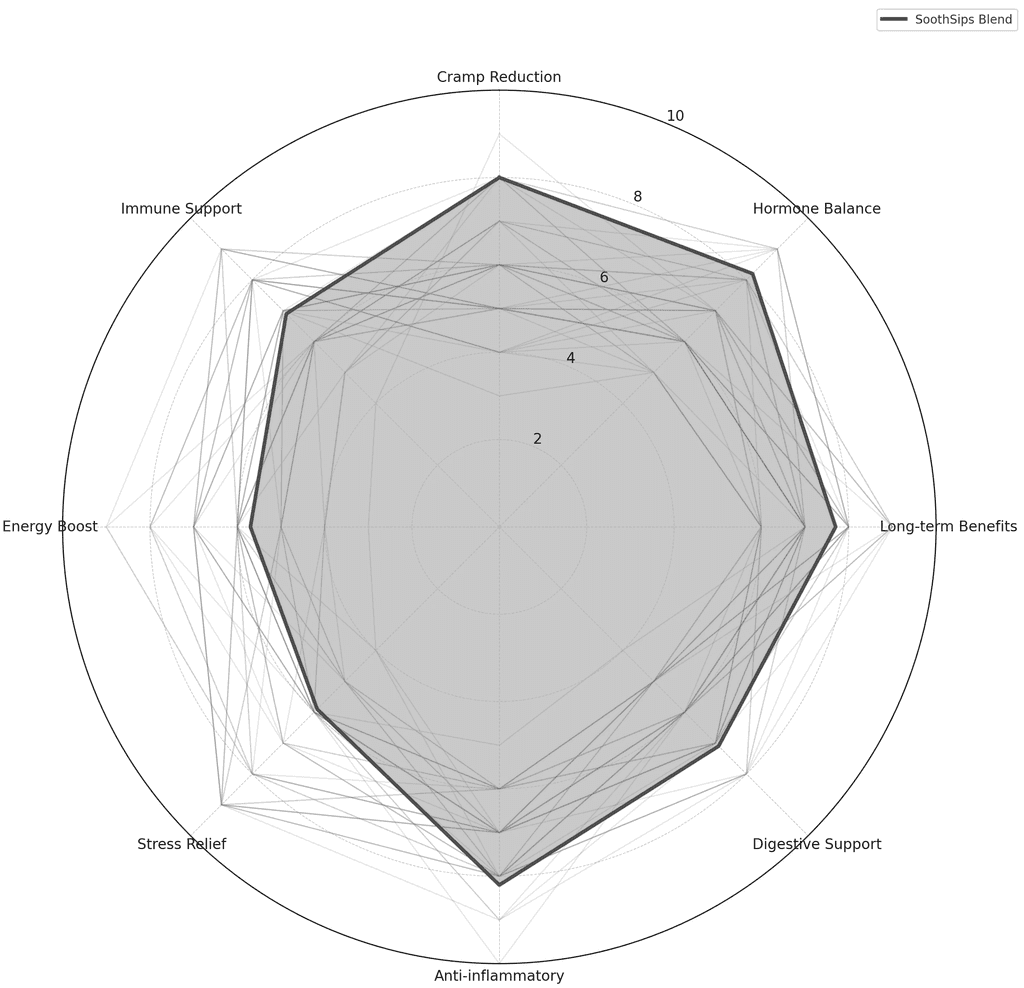
Formulation Optimisation
Multi-objective optimisation of compound ratios and preparation variables across high-dimensional interaction spaces, drawing on quantitative methods from portfolio theory.
04
Evaluation & Deployment
Structured validation of model outputs against reference datasets, with translation into interpretable, deployable decision systems for operational use.